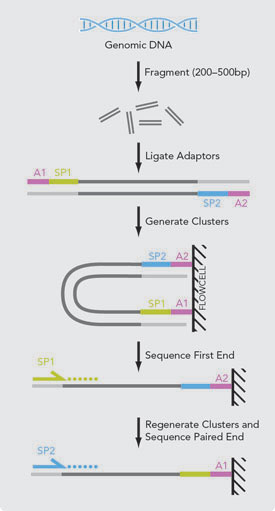
Illumina Sequencing | Illumina Sequencing by Synthesis – 1010Genome | Quality NGS Bioinformatics Data Analysis Services
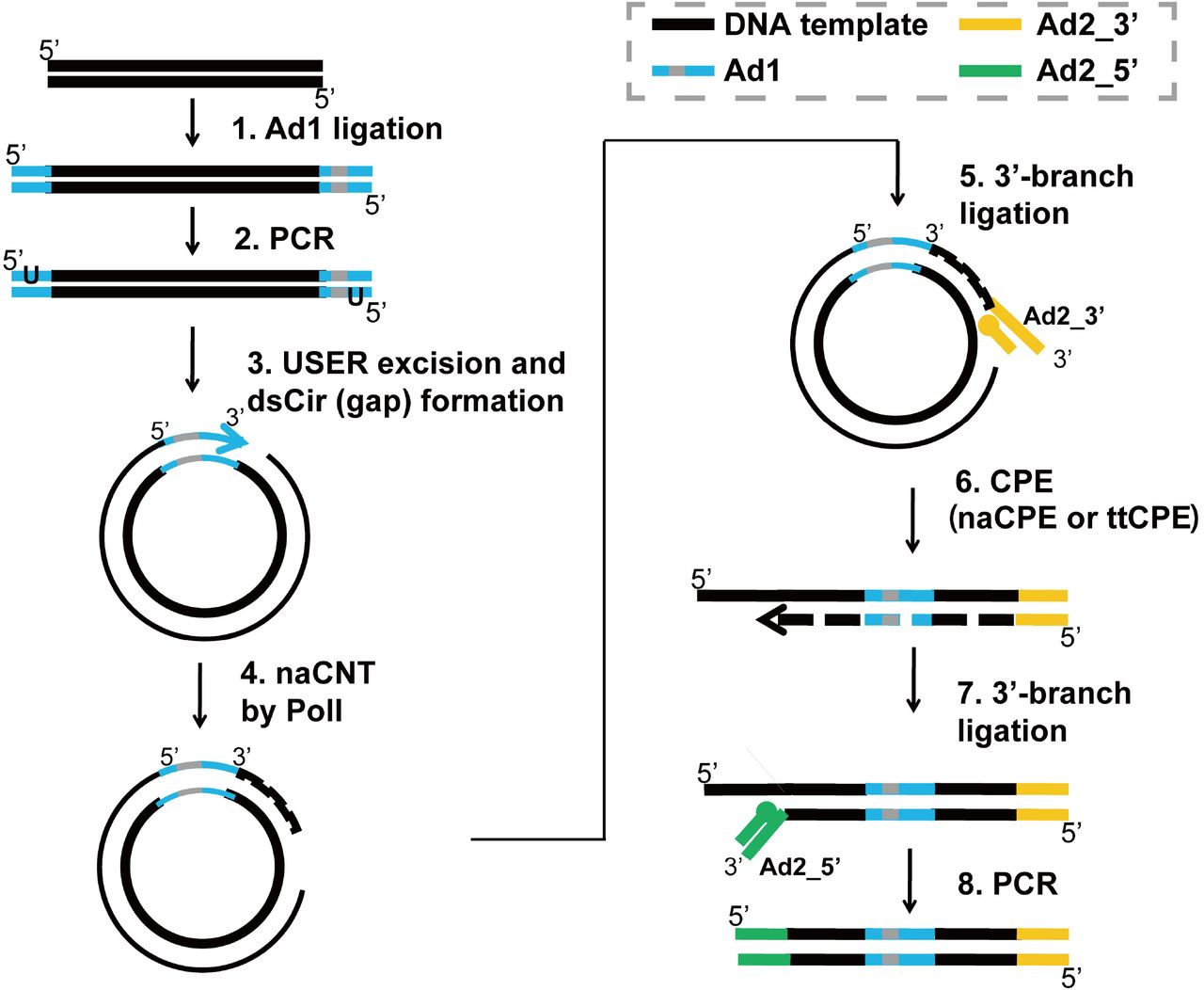
Development of Coupling Controlled Polymerizations by Adapter-ligation in Mate-pair Sequencing for Detection of Various Genomic Variants in One Single Assay | bioRxiv

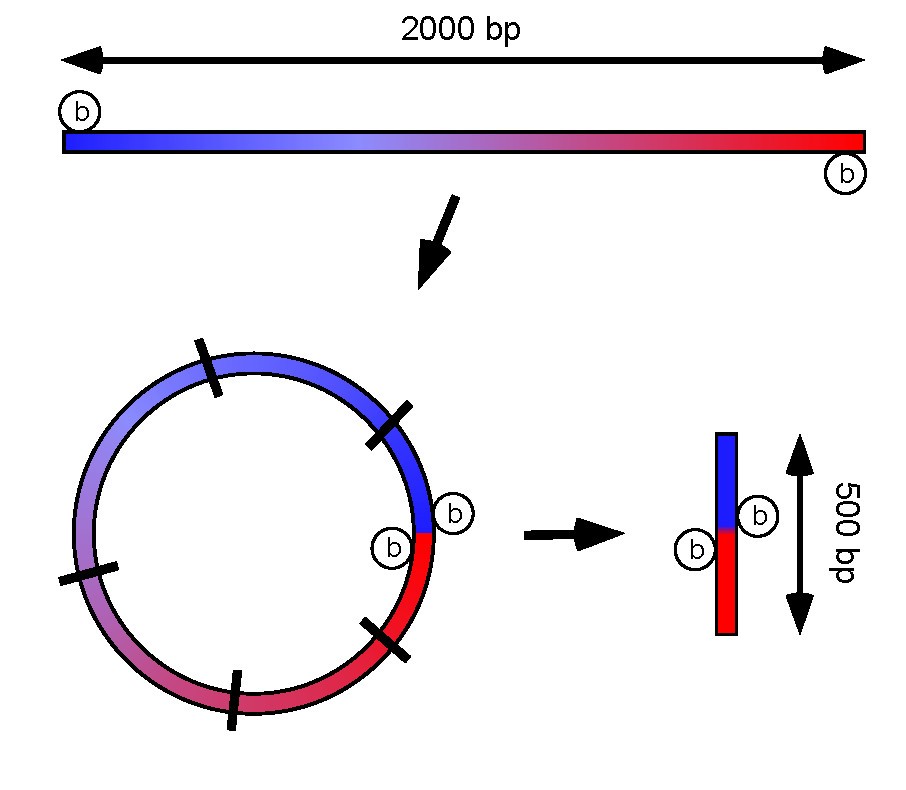
Conversion of Mate-Pair Reads into Long Sequences for Improving Assembly Scaffolding | Semantic Scholar

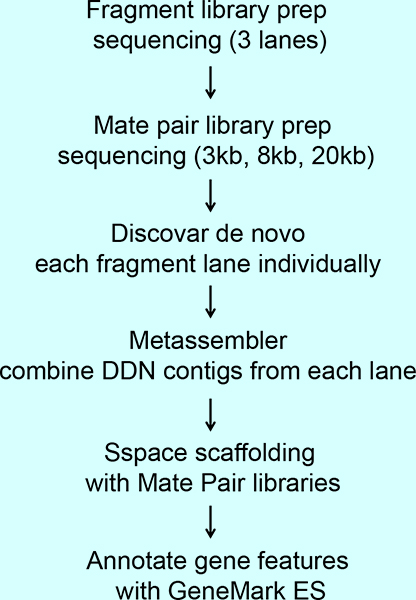
Optimization of the “in‐silico” mate‐pair method improves contiguity and accuracy of genome assembly - Zhou - 2023 - Ecology and Evolution - Wiley Online Library

Ion Torrent on X: "Ion TrueMate Library Kit technology generates mate-pair libraries with insert size of 2-8kb http://t.co/dkES1GZZv2 http://t.co/KpUjTRi2Ea" / X
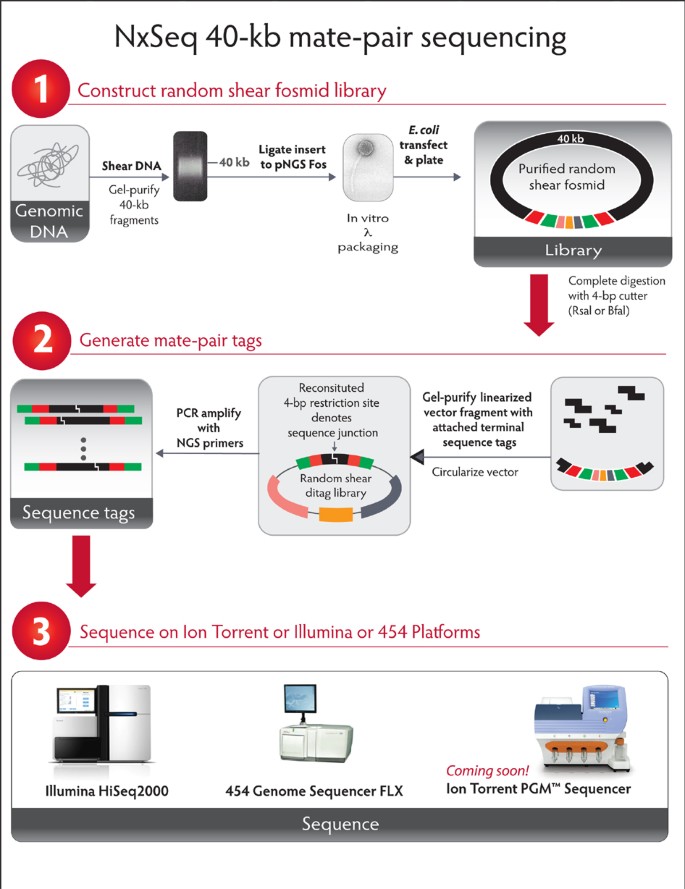
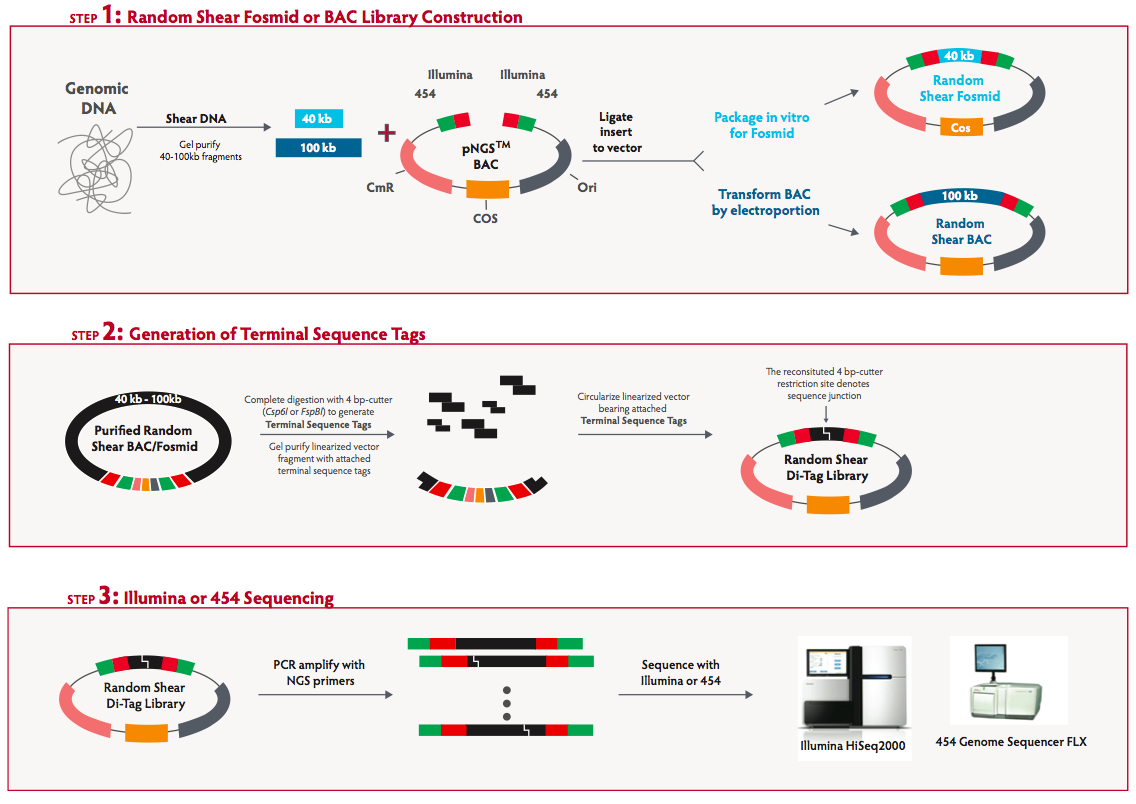
Long-span, mate-pair scaffolding and other methods for faster next-generation sequencing library creation | Nature Methods
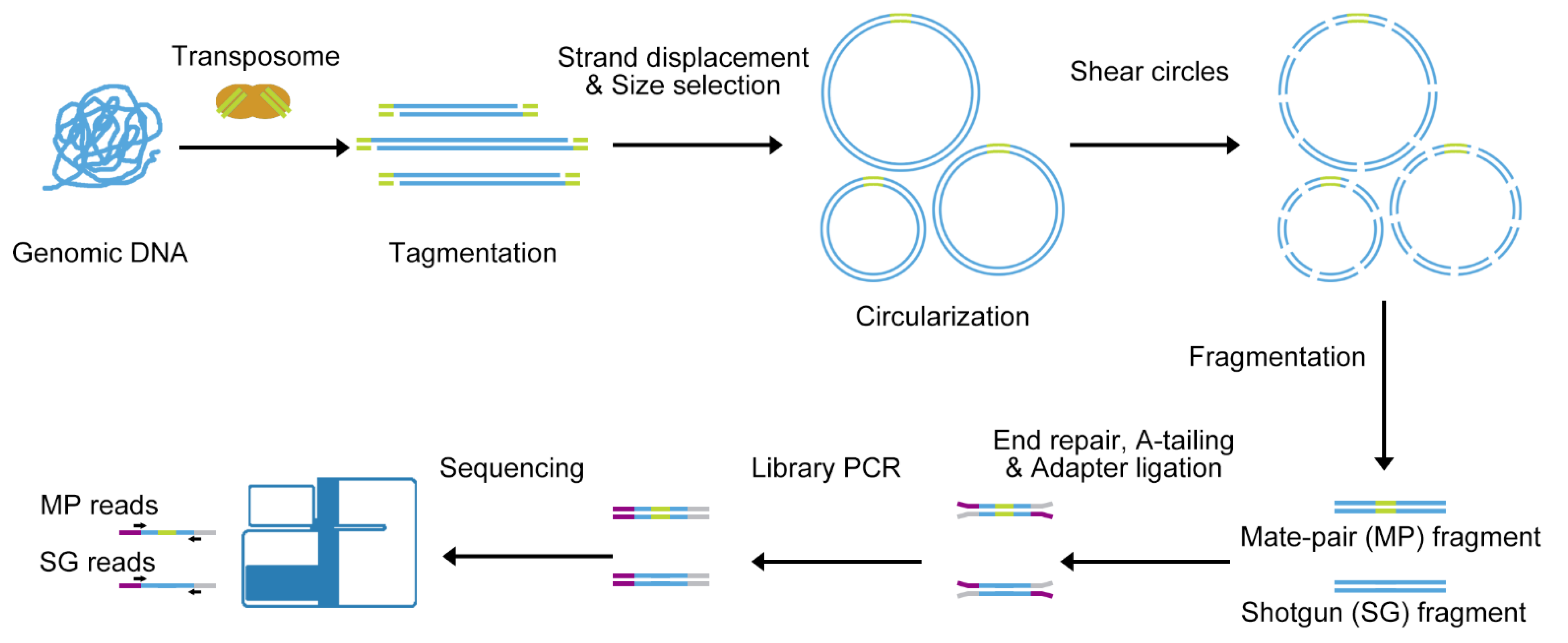

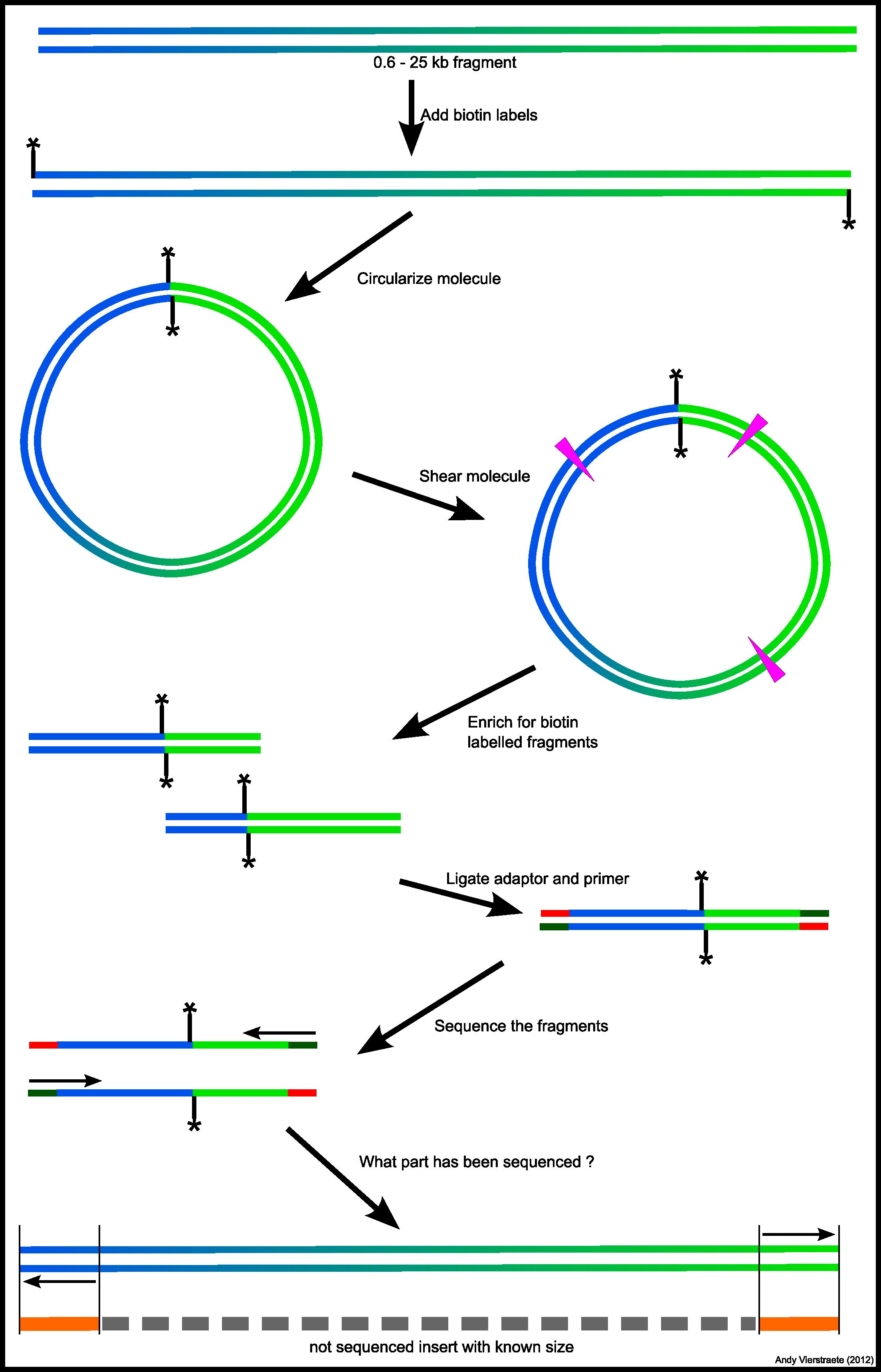
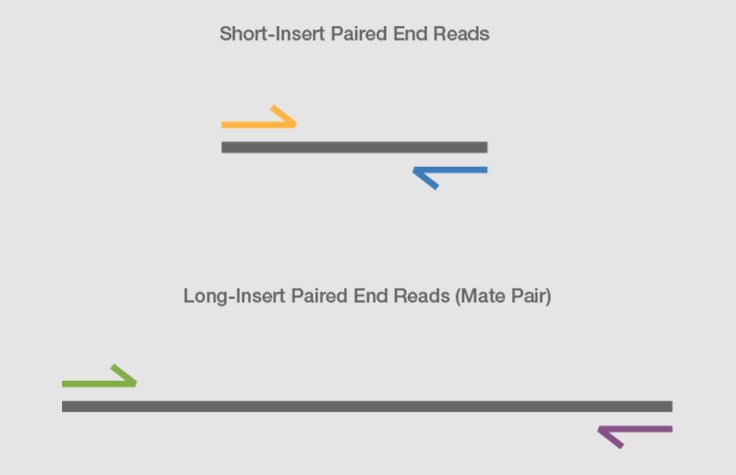
![PDF] Data Processing of Nextera Mate Pair Reads on Illumina Sequencing Platforms | Semantic Scholar PDF] Data Processing of Nextera Mate Pair Reads on Illumina Sequencing Platforms | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6979583dd405728ade1fce397fa6d102a941394c/2-Figure2-1.png)